bspmview
bspmview is a graphical user interface for overlaying, thresholding, and visualizing 3D statistical neuroimages in MATLAB. It currently requires a number of core functions from the MATLAB software [Statistical Parametric Mapping] (www.fil.ion.ucl.ac.uk/spm/), and indeed will work best with the statistical images produced using SPM (note that it has only been tested on SPM8/SPM12).
bspmview was directly inspired by and in some cases incorporates code from from two other statistical image viewers: xjview and FIVE. The idea behind bspmview was to integrate and improve some of the great features of these two viewing programs in an interface that I find to be more intuitive, accessible, and customizable. Most importantly, all of its features require no additional third-party software (other than SPM, of course). These features include:
reads .img/.hdr, .nii, and .nii.gz
can read non-statistical/non-intensity images (e.g., binary mask, ROIs, or count images)
quickly switch display of contrast sign (+, -, or +/-)
uncorrected and corrected image thresholding, including SPM-benchmarked familywise error rate (FWE) correction at both voxel and cluster levels
automatic anatomical labeling with the Anatomy Toolbox or Harvard Oxford atlases
generate interactive table of labeled activation peaks, and save (potentially) publication-ready tables to CSV
quickly generate beautiful and customizable surface renderings
save thresholded whole-brain maps or specific clusters as either intensity images or binary masks
save region-of-interest images by growing spheres/boxes around a coordinate (and optionally intersect the sphere/box with thresholded overlay image)
many colormap options
* and other random features, such using Neurosynth to examine functional connectivity and coactivation maps for the current coordinate
It requires a number of supporting utility functions and data files that should have been included in the distribution of bspmview. When bspmview is launched, it will look for these files in a folder called “supportfiles” that should be contained in the same folder as BSPMVIEW.
In addition, this software was inspired by and in some cases uses code from These are included in the “supporting files” folder that should have been included in the distribution of the main bspmview function.
Installation
- Download the software
- Put the bspmview folder either on your MATLAB path or in the “toolboxes” subfolder of your SPM installation. Note that only the parent folder needs to be on the path. The subfolder “supportfiles” is added to the MATLAB path whenever you run bspmview.
Starting
If you put bspmview in the “toolboxes” subfolder of your SPM installation, you will be able to launch bspmview by selecting it from the “Toolbox:” dropdown menu in the SPM graphical user interface. Otherwise, as long as bspmview is on your MATLAB path you can start it from the command line by entering:
>> bspmview
This will open a file selection dialogue window with which you can select the overlay image you’d like to display:
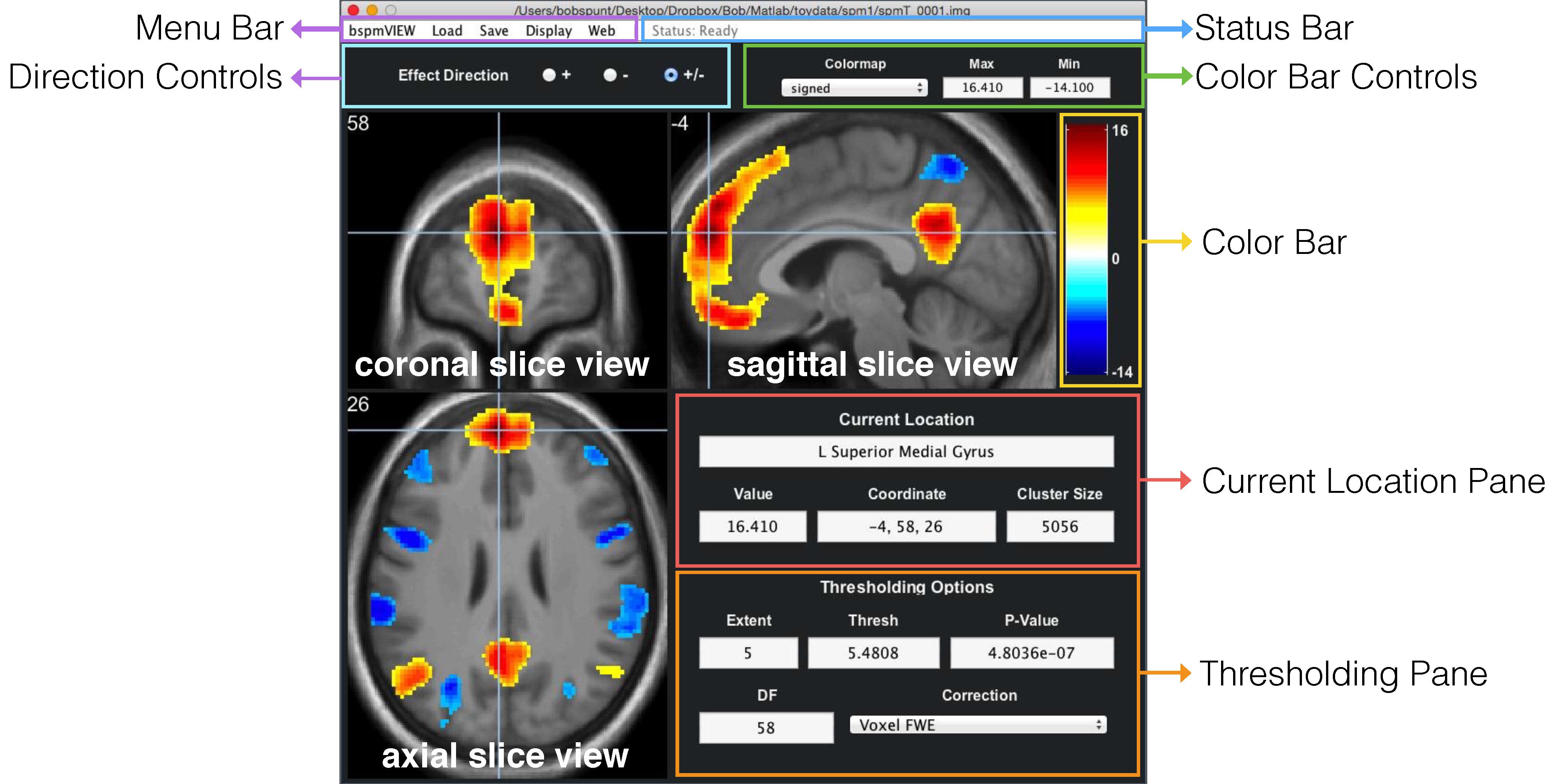
You can also give bspmview filenames for overlay and underlay images as arguments. The following will display the overlay spmT_0001.img on the default underlay:
>> bspmview('spmT_0001.img')
This will overlay “spmT_0001.img” on to “T1.nii”:
>> bspmview('spmT_0001.img', 'T1.nii')
Changing the Appearance of the Interface
So…
Modifying Display Preferences
So…
Creating a Surface Rendering
So…
Creating a Table of Results
So…
Have Questions?
Feel free to shoot me (Bob) an email at: bobspunt@gmail.com